e) Droplet overloading boosts the percentage of droplets filled with nuclei from 16.4% (obtained for the maximum loading concentration of the standard Chromium protocol) to 95.5% (obtained for 100-fold overloading using 1.53 million nuclei per channel). d) Representative images of droplets containing between one and ten nuclei, showing the overloading of a standard microfluidic droplet generator (10x Genomics Chromium). c) Detailed method design of scifi-RNA-seq. 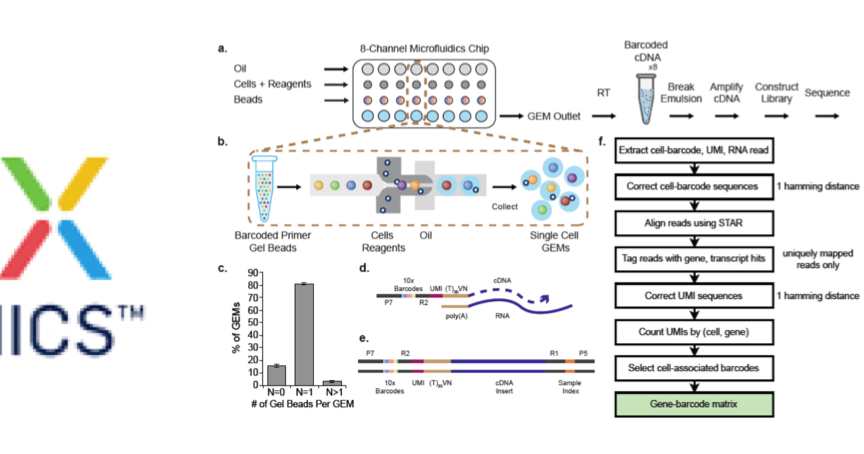
b) scifi-RNA-seq, which uses preindexing and droplet overloading to boost the throughput of droplet-based scRNA-seq. Scifi-RNA-seq combines preindexing of whole transcriptomes with droplet-based scRNA-seqĪ) Standard droplet-based scRNA-seq, where cells are loaded at a low concentration (limiting dilution) to avoid cell doublets, and most droplets do not receive a cell. Once the transcripts have been indexed inside the microfluidic droplets, the emulsion is broken, and the remaining steps of the library preparation are performed in bulk. All primers on a given bead carry the same barcode, which uniquely identifies the coencapsulated cell. Cells are then lysed inside the droplets, RNA molecules anneal to bead-tethered primers, and reverse transcription is performed inside the droplets. When aqueous and oil phases are combined at controlled flow rates, emulsion droplets coencapsulate individual cells with individual microbeads. In a typical droplet-based scRNA-seq experiment, the single-cell suspension is processed on a microfluidic chip, together with uniquely barcoded microbeads, reverse transcription reagents, and carrier oil ( Fig. With their high throughput, straight forward handling and consistent data quality, they have contributed to the wide adoption of single-cell RNA-seq (scRNA-seq) in many areas of basic biology and biomedical research. Microfluidic droplet generators are currently the most popular technology platform for single-cell sequencing 1– 3. We benchmarked scifi-RNA-seq on various human and mouse cell lines, validated it for primary human T cells, and applied it in a highly multiplexed CRISPR screen with single-cell transcriptome readout of T cell receptor activation. Compared to cell hashing methods, which flag and discard droplets containing more than one cell, scifi-RNA-seq resolves and retains individual transcriptomes from overloaded droplets. Compared with multiround combinatorial indexing, scifi-RNA-seq provides an easy and efficient workflow. Thereby, scifi-RNA-seq massively increases the throughput of droplet-based single-cell RNA-seq, and provides a straight forward way of multiplexing thousands of samples in a single experiment. Preindexing allows us to load several cells per droplet and computationally demultiplex their individual expression profiles. The scifi-RNA-seq assay combines one-step combinatorial preindexing of entire transcriptomes inside permeabilized cells with subsequent single-cell RNA-seq using microfluidics. To enable cost-effective single-cell sequencing for millions of individual cells, we developed ‘single-cell combinatorial fluidic indexing’ (scifi). Cell atlas projects and high-throughput perturbation screens require single-cell sequencing at a scale that is challenging with current technology.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
